ACTIVE Network API Developer Blog
The Prediction API: An Ensemble
Last week I posted an intro to our new Prediction API, with a simplified walk-through in Python, starting
from a set of asset descriptions labeled with topics and ending up with a topic prediction web service that accepts text and outputs topics and probabilities.
This week we're going to extend that example by adding a few more predictive models and a method to combine them into a stronger overall prediction that can capture
variability in the algorithms and the data. Since this isn't really a math, stats, or machine learning blog, I'll give quick overviews of the algorithms
and link to more details in footnotes. [tldr]
Building a Predictive Model
There are three basic steps to building a predictive model:
- Extract and process features from your data. For these examples, this means pulling out n-grams from our asset descriptions and vectorizing them. I'll post more about various methods for this later.
- Separate out training data to cover all your labels and a portion for testing. Here, our labels are the asset topics. There are many ways to do this, but for simplicity we just set aside a random portion of all the data.
- Run the training data through the algorithm many times with varying parameters and predict labels with your test data, assessing the model with some statistics or metrics. Repeat until your model performs to your liking.
Multinomial Naive Bayes (NB) Classifier
The naive Bayes1 is a probabilistic model, so for every class (topic here) we have, the model gives a percent likelihood that some data should be labeled with that class. That likelihood is based on a given set of features we extract from the data. In the last post,
we extracted n-grams and created a vector of n-gram counts and used them as features. The inverse concept is that each feature has a particular probability of being part of class. NB makes use of these relationships to learn the overall probabilities of each feature for each class.
NB makes some assumptions, including that the probability of a feature for a particular class is independent of the probabilities of the other features for that or any other class. This is counter-intuitive. After all, wouldn't the co-occurrence of both "mud" and "run" be a better indicator of a "mud run", and isn't it unlikely that "mud" would occur without "run" in this context? Well, we capture this in part by taking n-grams of 1, 2, and 3 tokens, but the assumption of independence actually proves to be less of an issue than you might think. More importantly, it means we don't need to calculate the cross-correlation of all the features and that saves lots of processing time.
NB makes some assumptions, including that the probability of a feature for a particular class is independent of the probabilities of the other features for that or any other class. This is counter-intuitive. After all, wouldn't the co-occurrence of both "mud" and "run" be a better indicator of a "mud run", and isn't it unlikely that "mud" would occur without "run" in this context? Well, we capture this in part by taking n-grams of 1, 2, and 3 tokens, but the assumption of independence actually proves to be less of an issue than you might think. More importantly, it means we don't need to calculate the cross-correlation of all the features and that saves lots of processing time.
The Support Vector Machine (SVM)
Support vector machines2 are popular and very powerful methods for classification. The general idea is to draw some sort of separating line between populations of data which
maximizes their separation, given a set of constraints, by maximizing the "margin" — the distance of that separator from the nearest training data point. The data can be mapped to higher dimensions using the "kernel trick" to achieve better separation:
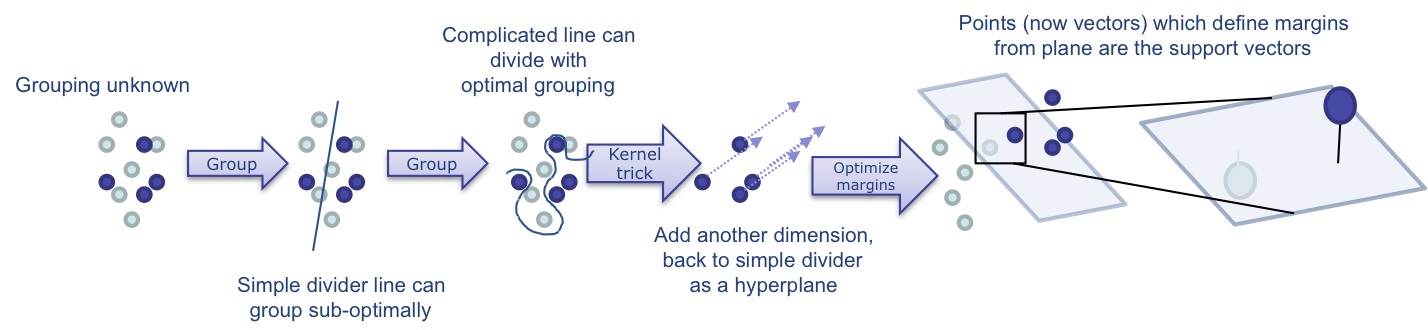
Of course, there are many ways to define and draw a line, but the fastest is a straight line and the linear SVMs are quite fast. By altering variables in the algorithm, you can adjust how this line is drawn, favoring things like good overall separation (large margin) over classification mistakes in the training data (error rate). Adjusting this trade-off is called "regularization", and amounts to adjusting the size and complexity of how the margin is represented. You may still want the best separating line even if some data points are on the wrong side or in the space between. However, as you test lines, you may want to penalize these data points, and you may want to do that differently depending on what the error was. This is done with so called "loss" functions. Varying regularization and loss functions affects results, but also processing time (both in training and in testing, and not necessarily the same way).

Of course, there are many ways to define and draw a line, but the fastest is a straight line and the linear SVMs are quite fast. By altering variables in the algorithm, you can adjust how this line is drawn, favoring things like good overall separation (large margin) over classification mistakes in the training data (error rate). Adjusting this trade-off is called "regularization", and amounts to adjusting the size and complexity of how the margin is represented. You may still want the best separating line even if some data points are on the wrong side or in the space between. However, as you test lines, you may want to penalize these data points, and you may want to do that differently depending on what the error was. This is done with so called "loss" functions. Varying regularization and loss functions affects results, but also processing time (both in training and in testing, and not necessarily the same way).
While searching for a solution to a problem like a line to separate data points, a common method is to sample some spot in the data, draw a line, test the separation, and then
move intelligently to another spot. Doing this repeatedly over all possible spots would eventually find the best separator, but in large data sets it would take forever. The gradient descent
method takes graded steps based on the comparisons during the testing and can eventually make assumptions about spots in the data it didn't visit based on spots it did visit. However, this
can still be time-consuming. Another variation called stochastic gradient descent3 tries to estimate areas of the data by repeatedly taking random samples of
the data and descending, building up an approximation of the data and the best solution. This process can adjust itself via a learning rate variable and has proven very efficient for large data sets, compared
to normal gradient descent. One nice side result of needing sub-samples of the data is that the learning can be done online, meaning you can learn or update the classification as the data comes in.
When you add functions to regularize or compute loss, you are also adding points of variability where you can decide to do more. The Passive-Agressive4 algorithm is one such
modifier which acts passively for data points with no loss according to some loss function, and aggressively with data points causing loss, to varying extents, to modify the loss function.
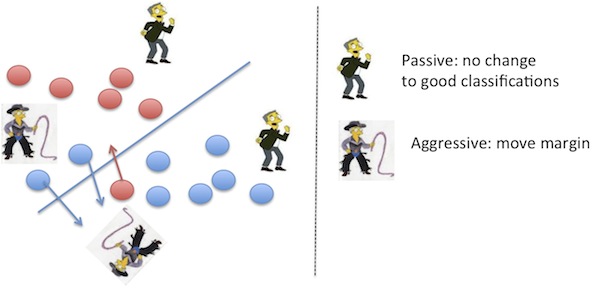
This guarantees that there is some non-zero margin at each update and means it can also be performed online. This was an addition Google made to weed out search result spam (2004-2005) and reduced it by 50%.

This guarantees that there is some non-zero margin at each update and means it can also be performed online. This was an addition Google made to weed out search result spam (2004-2005) and reduced it by 50%.
Scoring
Each of these methods has some level of internal scoring used to compare across the data and labels. Typically, to create a classifier out of these algorithms we extract that score and
call the best performer the winner. For probabilistic models this is the label with the highest probability, and for SVM this is the label furthest from the separating line (largest margin distance). This is fine for
comparing multiple runs on the same algorithm and getting the prediction. However, how do these scores relate to each other, and how can they be compared across algorithms? For example,
if an SVM algorithm for a particular prediction suggests "Swimming" with a distance of 6.12 and a Bayes algorithm suggests "Triathlon" with a probability of 0.85, which one should we put the most
confidence in? What do we do with negative distances or zero probabilities? Unfortunately, there are several ways to do this and none of them are consensus.
There are also several metrics used to assess the overall performance of a classifier. The typical metrics are precision and recall, and we can combine these two in what is called the F1 score. To account for varying label representation ("support", or
how many examples of a label you have in your training data), the F1 score can be weighted by the support for each label.
Ensembles
Combining models like this is generally termed an "ensemble" method. To accomplish this in our Prediction API, we do a few things:
- Normalize the distances in the SVM models (convert them to a uniform set of percentages)
- Adjust the normalized distance by internal probabilities of being on one side or the other of the margin to help compare
- Adjust probabilities or distances by the per-label weighted F1 score achieved on testing data for each model
- Average these resulting scores over all the models to get an overall top prediction.
The "adjust" step can take several forms, and other things can be done, like calibrating the probabilities to the actual distribution of labels in the test set, or track distances throughout the
training process for correlation normalization across all labels, etc. These are all things you can play with once you get the basics running smoothly.
So now let's build these...
We'll use the probabilistic Naive Bayes method good for this type of data, with sklearn's MultinomialNB predictor, as well as
two variations of the linear SVM from sklearn: SGDClassifier and
PassiveAgressiveClassifier.
In [5]:
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import SGDClassifier, PassiveAggressiveClassifier
models = []
models.append(MultinomialNB(alpha=.01))
models.append(SGDClassifier(alpha=.0001, loss='modified_huber',n_iter=50,n_jobs=-1,random_state=42,penalty='l2'))
models.append(PassiveAggressiveClassifier(C=1,n_iter=50,n_jobs=-1,random_state=42))
Save off the built models for later use, as well as some stats and metrics about the performance:
In [6]:
from sklearn.externals import joblib
from sklearn import metrics
results = {}
for model in models:
model_name = type(model).__name__
model.fit(X_train, Y_train)
joblib.dump(model, "models/topics."+model_name+".pkl")
pred = model.predict(X_test)
class_f1 = metrics.f1_score(Y_test, pred, average=None)
model_f1 = metrics.f1_score(Y_test, pred)
model_acc = metrics.accuracy_score(Y_test, pred)
probs = None
norm = False
try:
probs = model.predict_proba(X_test)
except:
pass
if probs is None:
try:
probs = model.decision_function(X_test)
norm = True
except:
print "Unable to extract probabilities"
pass
pass
results[model_name] = { "pred": pred,
"probs": probs,
"norm": norm,
"class_f1": class_f1,
"f1": model_f1,
"accuracy": model_acc }
results
Out[6]:
Let's define the scoring functions: compute_confidence performs the first portion to convert distances to a measure of confidence (or use probabilities if available), and then format
them. normalize_distances does the conversion we need for SVM. predict does the prediction and computes the confidence.
In [7]:
def normalize_distances(probs):
# get the probabilities of a positive and negative distance
pPos = float(len(np.where(np.array(probs) > 0)[0])) / float(len(probs))
pNeg = float(len(np.where(np.array(probs) < 0)[0])) / float(len(probs))
if pNeg > 0:
probs[np.where(probs > 0)] *= pNeg
if pPos > 0:
probs[np.where(probs < 0)] *= pPos
# subtract the mean and divide by standard deviation if non-zero
probs_std = np.std(probs, ddof=1)
if probs_std != 0:
probs = (probs - np.mean(probs)) / probs_std
# divide by the range if non-zero
pDiff = np.abs(np.max(probs) - np.min(probs))
if pDiff != 0:
probs /= pDiff
return probs
def compute_confidence(probs, labels, norm=False):
confidences = None
class_confidence = []
if norm:
confidences = normalize_distances(probs)
else:
confidences = probs
for c in xrange(len(confidences)):
confidence = round(confidences[c], 2)
if confidence != 0:
class_confidence.append({ "label": labels[c], "confidence": confidence })
return sorted(class_confidence, key=lambda k: k["confidence"], reverse=True)
def predict(txt, model, model_name, vectorizer, labels):
predObj = {}
X_data = vectorizer.transform([txt])
pred = model.predict(X_data)
label_id = int(pred[0])
predObj["model"] = model_name
predObj["label_name"] = labels[label_id]
predObj["f1_score"] = float(results[model_name]["class_f1"][label_id])
if results[model_name]["norm"]:
probs = model.decision_function(X_data)[0]
else:
probs = model.predict_proba(X_data)[0]
predObj["confidence"] = compute_confidence(probs, labels, norm=results[model_name]["norm"])
return predObj
To do ensemble prediction, first we run the predict method for each model:
In [8]:
prediction_results = []
label_scores = None
model_names = []
txt = "This is a tough mud run. Tough, as in, this could be one of the hardest events you, as a runner have ever attempted. This is not a walk in the park. This is not your average neighborhood 5k. This is more fun than a marathon. This is a challenge. The obstacles were designed by military and fitness experts and will test you to the max. Push personal limits while running, crawling, climbing, jumping, dragging, and other surprise tasks that test endurance and strength. The race consists of non-competitive heats as well as a free kid's course for children age 5-13. Each heat will depart in a specific wave so the course doesn't get overcrowded."
for model in models:
model_name = type(model).__name__
try:
prediction_results.append( predict(txt,
model,model_name, vectorizer, unique_topics))
model_names.append(model_name)
except Exception, e:
print "Error predicting" + str(e)
pass
prediction_results
Out[8]:
Finally, we do the weighted F1 adjustment:
In [9]:
from operator import itemgetter
conf_labels = {}
scored_labels = {}
for prediction_result in prediction_results:
if "label_name" in prediction_result:
l_name = prediction_result["label_name"]
f1_score = prediction_result["f1_score"]
conf = [ float(c["confidence"]) for c in prediction_result["confidence"] if c["label"] == prediction_result["label_name"] ]
if len(conf):
conf = conf[0]
else:
conf = 0
# update with confidence weighted by f1
#variations are to *= f1_score, or update confidence with just the confidence
if l_name in scored_labels.keys():
conf_labels[l_name] += f1_score * conf
scored_labels[l_name] += f1_score
else:
conf_labels[l_name] = f1_score * conf
scored_labels[l_name] = f1_score
labels_sorted = sorted(scored_labels.iteritems(),key=itemgetter(1),reverse=True)
labels_pred = []
# compute ensemble averages
num_models = len(models)
for i in xrange(len(labels_sorted)):
label = labels_sorted[i][0]
label_avg_f1score = float("%0.3f" % (float(labels_sorted[i][1]) / num_models))
label_avg_conf = float("%0.3f" % (float(conf_labels[label]) / num_models))
label_score = float("%0.3f" % (label_avg_f1score * label_avg_conf))
labels_pred.append({ "label": label, "score": label_score })
labels_pred = sorted(labels_pred, key=lambda k: k["score"], reverse=True)
labels_pred
Out[9]:
So, there's a quick implementation of an ensemble method combining linear and probabilistic machine learning for text document classification. Next we'll go more into the web service and
see how we can build models that perform their prediction faster, and how we can serve up those predictions fast enough for an enterprise-level Prediction API.
References
- Wikipedia does pretty good with its Naive Bayes page and the references therein. scikit-learn has a good description, as well.
- The OpenCV SVM intro page is pretty clear. scikit-learn explains SVMs, too.
- Wikipedia is concise and clear on SGD. scikit-learn's page is here.
- The canonical reference for online PA.
*Code here runs on Python 2.7.3 64-bit, with numpy 1.6.1, scipy 0.12.0, and sklearn 0.13.1. Ran/formatted with IPython 1.1.0 notepad.

0 Comments
Please sign in to post a comment.